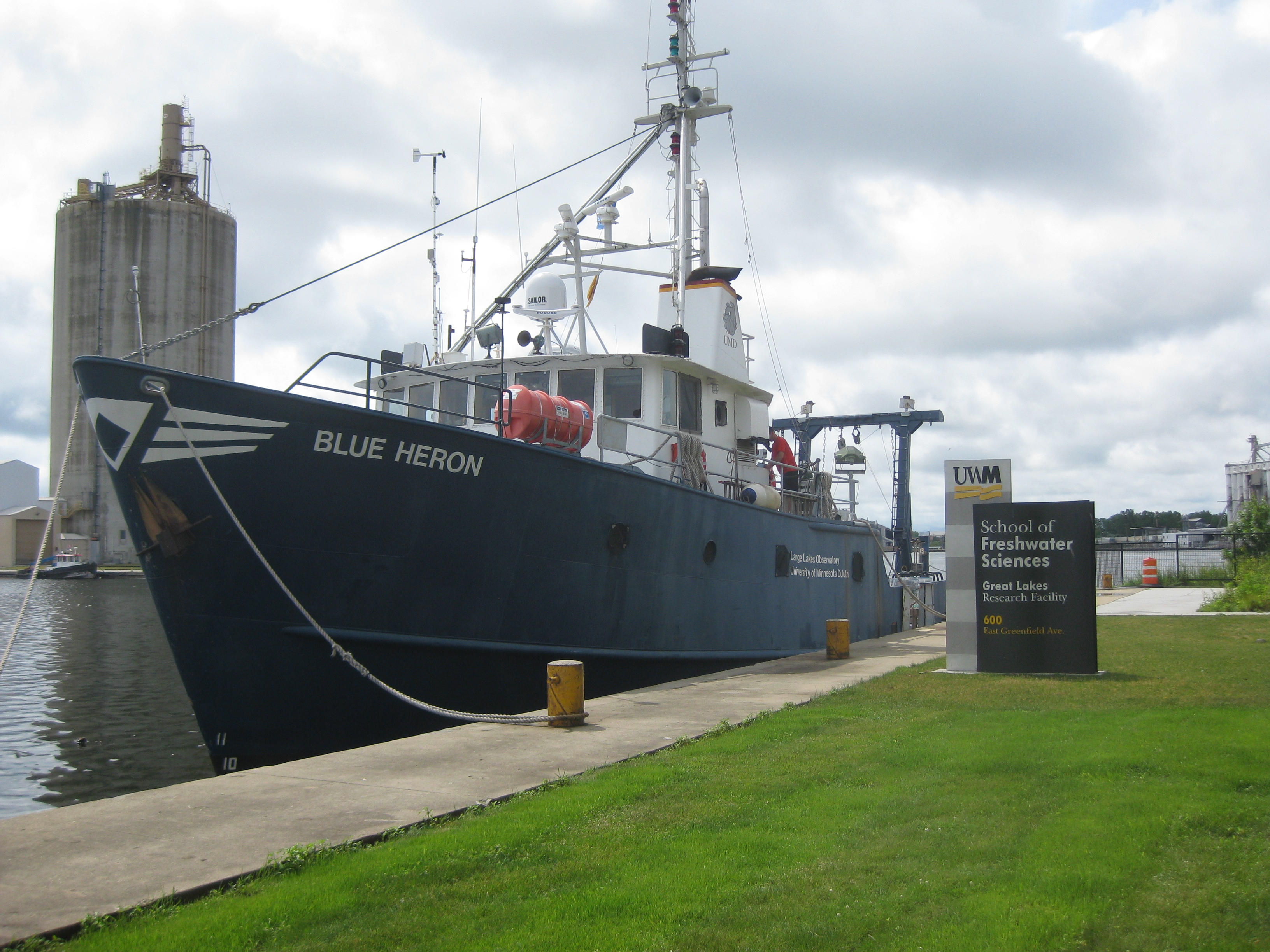
My summer research students Rebecca Lightman, Miranda Seixas, and I had the pleasure of sampling onboard the R/V Folger, with extra help from the Allen Lab. We were collecting samples for a new research project where we hope to investigate harmful cyanobacterial bloom dynamics in Lake Champlain.

Eggleston Lab Summer 2018 (L to R Miranda Seixas, Rebecca Lightman, Prof Eggleston)
Harmful algal blooms caused by cyanobacteria (CHABs) produce toxins that pose a major public health threat globally. The genes coding for these toxins are predicted to be acquired through a variety of mechanisms, including horizontal gene transfer. Viruses play an important role in shaping cyanobacterial populations and the exchange of genetic material. Transduction, the process whereby viruses transfer DNA between bacteria, is one horizontal gene transfer mechanism that could play a role toxin gene mobility. It has been suggested that cyanophage (viruses of cyanobacteria) may impact harmful algal bloom dynamics, and they may also mediate toxin gene exchange between cyanobacterial species.

Prepping the CTD Rosette (Miranda Seixas, Rebecca Lightman, Harper Baldwin)

Collecting water samples from the CTD Rosette
Eutrophication has led to increased incidence of CHABs in Lake Champlain, VT, which serves as a freshwater drinking supply and recreational attraction. The degree to which cyanophage mobilize cyanotoxin genes has implications for public health, as well as CHAB monitoring and management. This study aims to characterize cyanobacterial, cyanotoxin, and cyanophage diversity and dynamics to inform these public health and management decision making processes. While cyanotoxin gene transduction by cyanophage has been predicted, this will be this first study to directly test this hypothesis.

Miranda and Rebecca fixing samples for microscopy.

Miranda preparing slides for microscopy while Rebecca filters water.
At Middlebury College we are very lucky to have access to this amazing research vessel, the R/V David Folger which allows us to gain access to sampling areas all around Lake Champlain. For this trip we cruised up to Shelburne Bay, with Captain Rich Furbush and crew member Chris Goodrich, as well as some crew trainees. We collected water using the CTD rosette with niskin bottles (a specialized set of water bottles, with probes that detect conductivity, temperature and depth). Once we had the water on deck it was portioned out for a variety of downstream applications. We filtered and stained small volumes of water to count bacteria and viruses in the water. Additional samples were collected on filters and stored for future work extracting and identifying bacterial communities to study their DNA. The final aspect of the sampling was to collect a large volume of water for viral precipitation, which we we also analyze DNA from, to determine all of the types of viruses present in the water sample. We hope to be able to focus our DNA analysis on cyanobacteria and cyanophage (their viruses) to better understand the role of viruses in the bloom dynamics.